 Additional cDNA is engineered, in the phage display vector, after the VL and VH sequences to allow characterization and purification of the mAb produced. In addition to the ability to produce phage displaying the mAb, the phage display vector can be used to produce the mAb itself (not attached to phage capsid proteins) in certain strains of E. The result is a library of phages, each expressing on its surface a mAb and harboring the vector with the respective nucleotide sequence within ( Figure 1c). coli that are transformed with the phage display vector library. For those genes, a helper phage is added to the E. However, the phage display vector pComb3X does not have all the other genes necessary to encode a full bacteriophage in E. 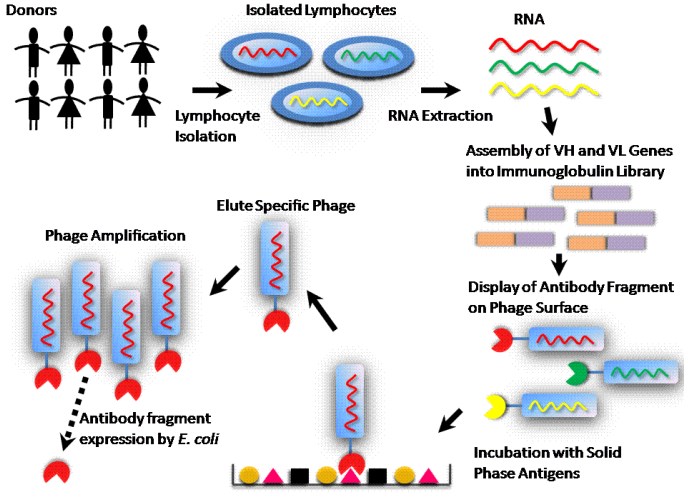
The VH and VL PCR products, representing the antibody repertoire, are ligated into a phage display vector (e.g., the phagemid pComb3X) that is engineered to express the VH and VL as an scFv fused to the pIII minor capsid protein of a filamentous bacteriophage of Escherichia coli that was originally derived from the M13 bacteriophage. Variations on phage display libraries include (i) libraries constructed for Ig isotypes (e.g., IgG, IgA, and IgE) and (ii) libraries of mAbs expressed as Fab fragments or as single-chain variable fragments (scFv), the latter of which consist of the VH and VL joined by a linker ( Figure 1b). This immortalizes recombinant cDNA clones for expressed Igs. Defined sets of primers specific for the different VH and VL chain–region gene families then allow amplification of all transcribed rearranged variable regions within a given immunoglobulin repertoire for library construction, thus reflecting all antibody specificities in a particular individual. This RNA is reverse-transcribed into cDNA, which is used for PCR of the VH and VL chains of the encoded antibodies ( Figure 1a, b). The key to success is preparation of quality RNA from the cell source chosen (e.g., peripheral blood mononuclear cells). A large antibody library and efficient selection are needed to isolate specific mAbs from a cloned immunoglobulin repertoire. The APD process begins with antibody-library preparation, followed by ligation of the variable heavy (VH) and variable light (VL) PCR products into a phage display vector, culminating in analysis of clones of mAbs. These characteristics make APD a powerful tool to better understand immunological processes and human diseases that involve formation of (auto-)antibodies against defined (self-)antigens. 
This technique allows in vitro selection of mAbs of virtually any specificity, greatly facilitating recombinant production of reagents for use in research and clinical diagnostics, as well as for pharmaceuticals for therapeutic use in humans (e.g., adalimumab, the first fully human APD-derived mAb) ( Lee et al., 2007).īecause of a physical connection established between the antibody fragment on the outside and the genetic information encoding the displayed protein within the phage, APD also allows comprehensive studies of genetics and function of antigen-specific mAbs. Antibody phage display (APD) is based on genetic engineering of bacteriophages (viruses that infect bacteria) and repeated rounds of antigen-guided selection and phage propagation ( Barbas, 2001).  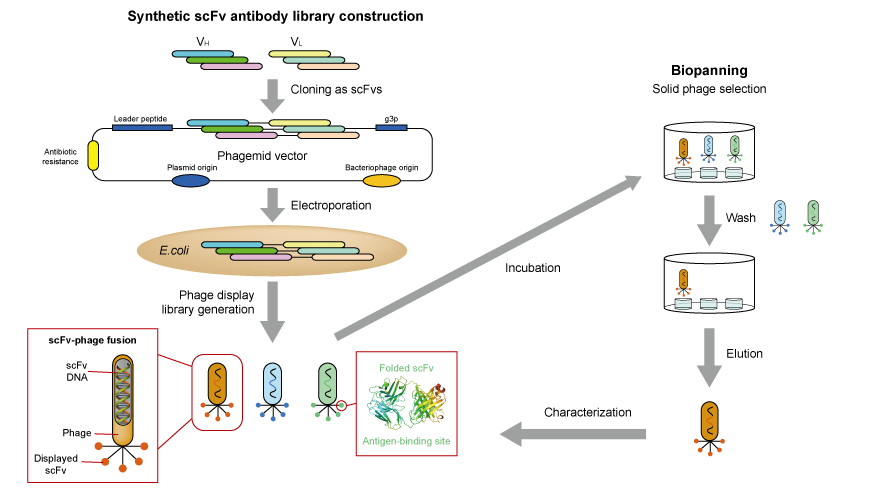
The production of human monoclonal mAbs for research and clinical use is closely related to the development of phage display technology, initially described by Smith in 1985 and further developed by other groups (e.g., Winter, McCafferty, Lerner, Barbas). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
